Introduction¶
Technological advances are revolutionizing methods in neuroscience. The vast collection of recording setups used, and the large variety of experimental subjects puts high demands on flexible data organization.
Often in neuroscience, the experimental setup is not finalized or rigidly predefined before data acquisition begins. Results may require additional branches of experimentation or reevaluation of the setup. Put simply, experiments have a tendency to organically grow along the experimental time line.
Expipe is thus introduced as an organizational tool that grow organically together with the experimentation - to ease data management in such experimental paradigms.
The aims of Expipe is to organize data and metadata in a way such that they:
are flexible towards a multitude of user aspects.
are readable for humans and machines for many years to come.
are sharable.
support high throughput data analysis.
support multiple types of large-scale data sets.
To this end we use the flexible filesystem as a non structured (NoSQL) type database to store data and metadata. This way of storing metadata consist of assigning key-value pairs as Python dictionaries.
An Expipe project contains the object types actions, entities, modules
and templates.
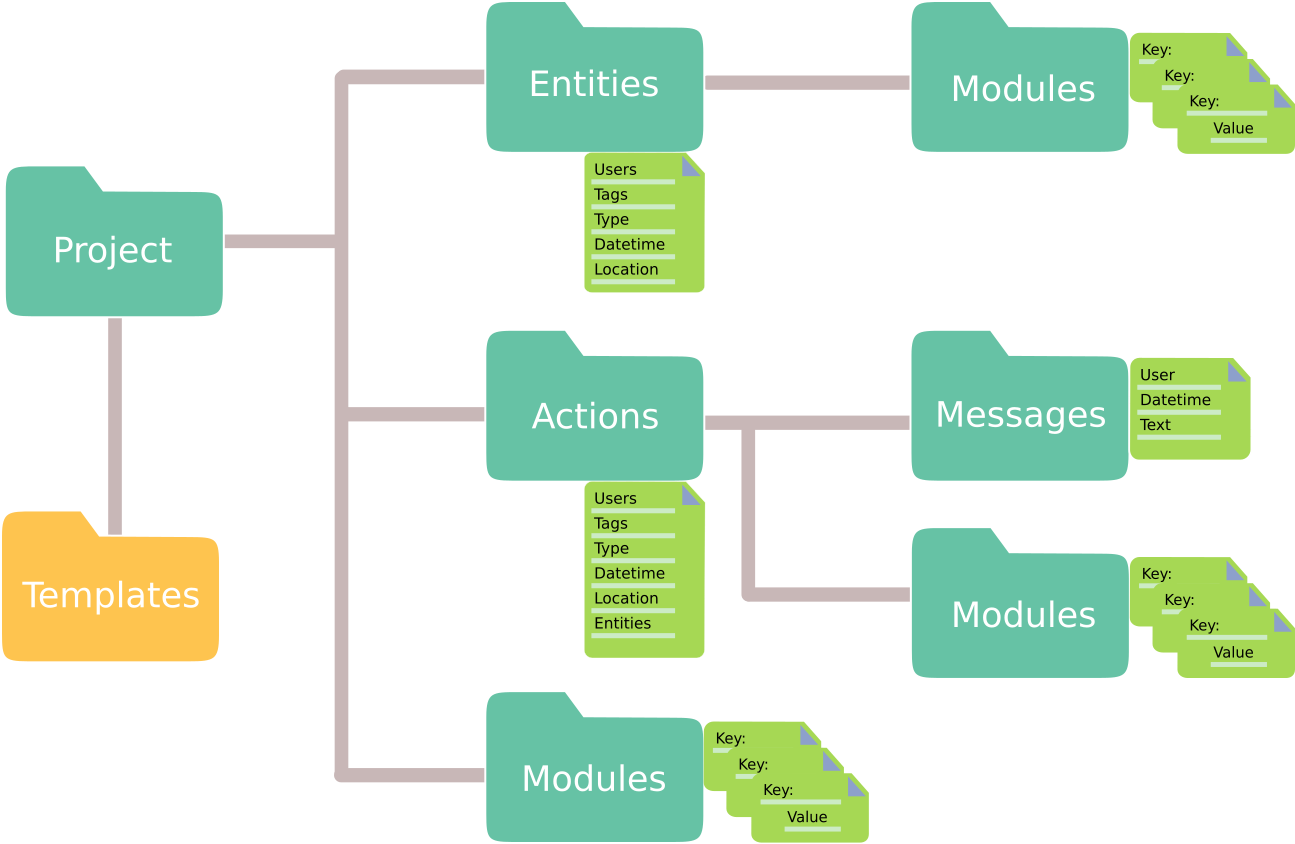
Together, these objects represent an experimental project at large.
modules sit at the core of the system and are used to describe projects, actions
and entities in detail.
The modules typically contain metadata about the equipment, environment, or subjects,
such as the numerical aperture of a microscope lens,
the serial number of an acquisition system,
or the temperature of a room.
actions define events that occurred at a specific time,
such as an experiment, an analysis, or a simulation.
actions have a few specific attributes, such as a timestamp, and store detailed metadata in modules.
Entities are long-lived things that are used in an action,
typically an experimental subject such as a rat or mouse.
actions refer to Entities, but but they do not link directly to them.
messages are user specific lines of text added to actions, such as notes.
During an experiment metadata can be automatically added by user specific
templates.
Templates are prefilled key-value pairs describing all aspects
of your experiments e.g. recording environment, acquisition system etc.
When added, templates are introduced as modules which are descriptors of
project and/or action entities. A project is the
root object during communication with expipe and contain modules and actions.
actions are individual, well actions, of interaction with experimental assets
or expipe itself such as recordings or analysis respectively.
actions also contain modules which are specific to a particular action in
contrast to project modules which are more general.
We encourage users from neuroscience to base templates on the odML terminologies which can be a priori filled out by a user or added in an empty state for a posteriori documentation.